Foldit
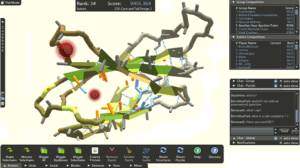
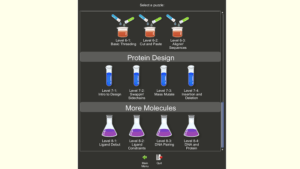
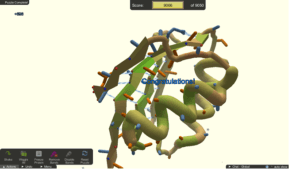
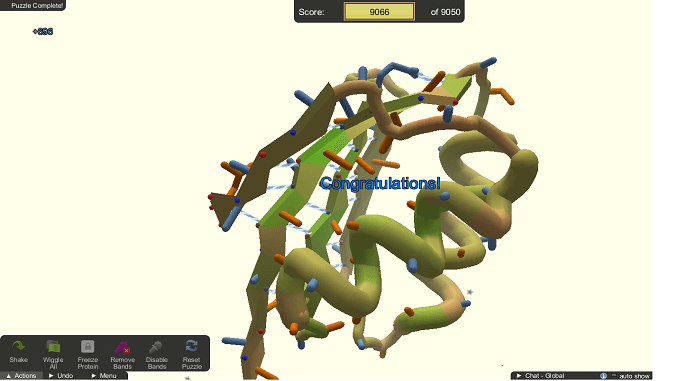
Knowing the structure of a protein is key to understanding how it works and to targeting it with drugs. The number of different ways even a small protein can fold is astronomical. Foldit attempts to predict the structure of a protein by taking advantage of humans’ puzzle-solving intuitions and having people play competitively and collaboratively to fold the best proteins in 3D puzzle games. Foldit is developed by the Center for Game Science at University of Washington in collaboration with UW Department of Biochemistry.
Articles
Computer gamers solve medical problems
As a core area of molecular biology, the study of protein structure is integral to elucidating pathological processes. Protein structure prediction has become one of the most important goals of pharmaceutical and biochemical industries. With […]

Scientific American – on Citizen Science Games
Can You Diagnose Dementia from a Gaming App? By Bahar Gholipour on November 18, 2016 SAN DIEGO—You are guiding a ship through rough waters. On your way you may encounter magical creatures. You can snap […]
Publications

Foldit players contribute to the CASP experiment
CASP is a worldwide experiment in which groups work to solve structures of unknown proteins. Many @Foldit players are listed as co-authors in this latest Nature paper examining the results of the WeFold "coopetition" during […]

Foldit Standalone: a video game-derived protein structure manipulation interface using Rosetta
Foldit Standalone is an interactive graphical interface to the Rosetta molecular modeling package. In contrast to most command-line or batch interactions with Rosetta, Foldit Standalone is designed to allow easy, real-time, direct manipulation of protein […]
Interviews

Radio interview on Science Friday, Adam Gazzaley and Zoran Popovic
Radio interview on Science Friday, a weekly radio show and website covering science, technology and other cool stuff. For the game Neuroracer: Adam Gazzaley, an associate professor of neurology, physiology and psychiatry at the University […]